Applications/ActinAnalyzer2D: Difference between revisions
| Line 95: | Line 95: | ||
===== Sample data ===== | ===== Sample data ===== | ||
For testing the ''ActinAnalyzer2D'' operator we provide some sample data: | |||
[http://www.informatik.uni-halle.de/mitobo/downloads/actin_examples.zip ActinAnalyzer2D sample data] | |||
The archive contains the following sub-folders: | |||
* ''imageData'': test images that were used in the ICPR publication mentioned above | |||
* ''maskData'': corresponding label images | |||
* ''featureData'': pre-calculated features for the cells (re-calculating the features may require up to an hour, depending on the machine used) | |||
* ''resultData'': sample results calculated on the given data | |||
In addition, in the archive a file with a sample parameter configuration for the operator can be found. The parameters are those used for producing the sample results. Once the operator has been started the file can be loaded via the '' 'File' '' menu and its entry '' 'Load Settings' ''. Note that you need to set the image and mask directories, and also the feature directory according to your local file system structure and the place to where you extracted the zip file. | |||
=== Updates === | === Updates === | ||
Revision as of 18:28, 30 July 2014
Actin Analyzer 2D
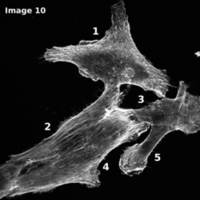

The Actin Analyzer 2D is available since release version 1.4 of MiToBo.
Related Publications
- B. Möller, E. Piltz and N. Bley, "Quantification of Actin Structures using Unsupervised Pattern Analysis Techniques".
In Proc. of Int. Conf. on Pattern Recognition (ICPR '14), IEEE, Stockholm, Sweden, August 2014, accepted for publication.
Name of Plugin/Operator
de.unihalle.informatik.MiToBo.apps.actinAnalysis.ActinAnalyzer2D
(available since MiToBo version 1.4)
Main features
- automatic extraction of different structural patterns by unsupervised texture analysis and clustering
- co-occurence matrices and Haralick features are currently used for texture charakterization
- structure quantification performed based on cell-wise cluster distributions
Usage
To run the ActinAnalyzer2D perform the following steps:
- install MiToBo by following the instructions on the Installation page
- run MiToBo and start the operator runner by selecting the menu item MiToBo Runner from Plugins -> MiToBo
- in the selection menu navigate to 'de.unihalle.informatik.MiToBo.apps.actinAnalysis' and select the operator ActinAnalyzer2D
This will bring up the operator window of the ActinAnalyzer2D.
- Input data:
The operator analyzes all images in the given input folder, expecting images to have the extension ".tif".
If the images contain more than one channel, the first channel is used. In addition, for each image
either a label image of pre-segmented cell regions or a set of region boundaries is expected to be available in the given mask folder.
These files should have the same basenames like the corresponding images, but either end on "-mask.tif" in case of label images,
or on ".zip" or ".roi", respectively, in case of region sets. Note that the operator currently only accepts ImageJ 1.x ROI sets as input.
If label images are used as segmentation masks, the labels of individual cells in the label images are used as unique identifiers for the cells, i.e., are used as labels in the various plots. If the segmentation data is given in terms of ROIs a unique identifier is derived from the order of the cell boundaries in the ROI set. Note that in the latter case a mask image is written to the output directory as additional output where the cells are marked by their identifiers to allow for easier assessment of the results.
The operator is able to automatically consider different groups of cells in its analysis, e.g., generate plots of the cluster distributions for each group individually.
However, the group membership of each image has to be encoded in its filename. In detail, the operator expects the file names to obey the following structure:groupName_imageID.tif
In particular, there must not occur more than a single underscore in each filename and the imageID must be unique for each image of a group.
If the image names do not follow these requirements, all images are treated as a single group. - Output data:
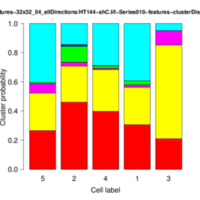
The operator displays the cluster distributions for each group of cells as stacked bar plots and box-whisker plots. In addition, it writes several files to the given output folder:- *-features.txt: feature data for each image
- *-features.tif: image stack visualizing the feature data
- *-features-config.ald: configuration of the operator in this run
- *-clusterDistro.txt: cluster distributions per image
- *-clusters.tif: pseudo-colored image illustrating the cluster distribution per image
- AllImagesClusterStatistics.txt: cluster distribution raw data for all images
- AllImagesSubspaceFeatures.txt: if PCA is applied to the cluster distributions prior to the distance calculations, this file contains the subspace feature vectors
- AllImagesPairwiseDistanceData.txt: matrix of pairwise Euclidean distances for distribution vectors, can be examined, e.g., with Multidendrograms (see below)
- *-distribution.png: for each cell group a stacked bar plot is saved showing the cluster distribution for each cell of the group
- Parameters:
| Name | Description |
| Image directory | directory where the input image data can be found |
| Mask directory | directory where the label images or region boundary files can be found |
| Mask format | format of the segmentation data files: LABEL_IMAGE = images with unique labels for each cell and a value of zero for the background / IJ_ROIS = set of ImageJ 1.x ROIs, one ROI for each cell |
| Output and working directory | directory to which the result files and intermediate data is written |
| Calculate features | if disabled the operator expects the features to be already present in the input directory and skips the (time-consuming) feature calculations; this option is helpful if the features have already been calculated ones and only the parameters of the clustering should be changed |
| Feature directory | directory where the features should be saved or - in case they are already available - from where they are read; the directory can be the same as the output and working directory |
| Tile size x/y | size of the sliding window used for feature calculations, should be chosen according to the resolution of the input images |
| Tile shift x/y | shift of the sliding window, if the shift is smaller than the tile size sliding windows overlap |
| Distance | pixel-pair distance in co-occurence matrix calculations |
| Set of directions | directions to be considered in co-occurence matrix calculations |
| Isotropic calculations | the texture features are derived from co-occurence matrixes; if this flag is enabled features for different directions are averaged, otherwise all individual directions are preserved (resulting in larger, but also more informative feature vectors) |
| Number of feature clusters | number of clusters in first stage, i.e., number of expected structural patterns in the images |
| Do PCA in stage II? | allows to enable/disable PCA on the cluster distribution vectors prior to the pairwise distance calculations; by default enabled |
Additional Tools
The hierarchical clustering in stage II of our approach as described in the paper has been done using the MultiDendrograms software.
In principal every hierarchical clustering tool can be applied.
The basis for the hierarchical cluster analysis is the file AllImagesPairwiseDistanceData.txt to be found in the output directory upon termination of an analysis run. It contains a matrix of pairwise Euclidean distances of the (optionally dimension-reduced) cluster distribution vectors of all cells. The file can directly be loaded by MultiDendrograms, for other tools format convertion might be necessary.
You can download the latest version of MultiDendrograms from its webpage: [1]
Sample data
For testing the ActinAnalyzer2D operator we provide some sample data: ActinAnalyzer2D sample data
The archive contains the following sub-folders:
- imageData: test images that were used in the ICPR publication mentioned above
- maskData: corresponding label images
- featureData: pre-calculated features for the cells (re-calculating the features may require up to an hour, depending on the machine used)
- resultData: sample results calculated on the given data
In addition, in the archive a file with a sample parameter configuration for the operator can be found. The parameters are those used for producing the sample results. Once the operator has been started the file can be loaded via the 'File' menu and its entry 'Load Settings' . Note that you need to set the image and mask directories, and also the feature directory according to your local file system structure and the place to where you extracted the zip file.
Updates
July 2014
- Released first version of Actin Analyzer as published in ICPR 2014.