Applications/MTBCellCounter: Difference between revisions
(Created page with "<br/> <br/> == MTB Cell Counter == 150px|right|link= 150px|right|link= [[File:ActinDistro.png|150px|right|lin...") |
No edit summary |
||
| Line 60: | Line 60: | ||
** 'Remove': delete the last type from the end of the list | ** 'Remove': delete the last type from the end of the list | ||
** 'Delete': delete the last placed marker | ** 'Delete': delete the last placed marker | ||
** 'Reset': deletes all markers | |||
** 'Show Markers': enables/disables display of markers | |||
** 'Show Numbers': enables/disables display of marker numbers | |||
** 'Show All': enables/disables display of both markers and numbers | |||
* Results: | |||
** 'Results': shows table with marker statistics | |||
** 'Save Markers': saves the markers to an XML file | |||
** 'Load Markers': loads markers from an XML file | |||
** 'Export Image': save a copy of the image including all markers | |||
<br> | |||
For the pre-segmentation of spot-like structures a particle detector operator of MiToBo is used. The operator offers the following parameters: | |||
* Parameters: | * Parameters: | ||
{|class="wikitable" | {|class="wikitable" | ||
| Line 68: | Line 78: | ||
|Description | |Description | ||
|- | |- | ||
|'' | |''Minimal Scale (JMin)'' | ||
| | |scale of smallest particles, integer value >= 1 | ||
|- | |- | ||
|'' | |''Maximal Scale (JMax)'' | ||
| | |scale of largest particles, integer value > JMin | ||
|- | |- | ||
|'' | |''Scale Interval Size'' | ||
| | |several adjacent scales are correlated, the interval size determines how many scales are considered | ||
|- | |- | ||
|'' | |''Correlation Threshold'' | ||
| | |threshold for detecting particles, the smaller the threshold is chosen the more particles will be detected | ||
|- | |- | ||
|'' | |''Minimum Region Size'' | ||
| | |minimal size of valid particles, smaller particles are discarded | ||
|} | |} | ||
===== Sample data ===== | ===== Sample data ===== | ||
Will be provided soon... | |||
Revision as of 15:52, 11 March 2015
MTB Cell Counter
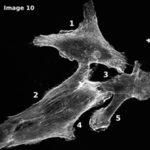

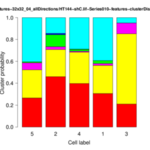
The MTB_CellCounter plugin is available since release version 1.5 of MiToBo.
The plugin is based upon the original Cell Counter plugin for ImageJ written by Kurt De Vos (Cell Counter) and now available in Fiji. Compared to the original version the MTBCellCounter plugin adds some nice new features:
- pre-segmentation and filtering of spot-like structures
- free configuration of marker colors
- advanced editing of markers
- status bar, tooltips and keyboard shortcuts
Related Publications
- The Authors,"The 'MTB Cell Counter' - a versatile tool for semiautomated quantification of sub-cellular phenotypes in fluo-
rescence microscopy images".
2015, submitted for publication.
Name of Plugin/Operator
mtb_cellcounter.MTB_CellCounter
(available since MiToBo version 1.5)
Usage
For using the plugin you need to install MiToBo by following the instructions on the Installation page. Running ImageJ you will then find a new entry 'MiToBo' in the plugins menu from where you can select the 'MTB CellCounter' plugin.
The usage of the plugin is leaned on the usage of the original plugin (see also Cell Counter webpage).
The basic workflow is as follows:
- open the image you would like to process and press the 'Initialize' button
- optionally configure the particle detector via the 'Configure operator...' button and then press 'Detect'
- once the detection is finished you can filter detected particles via the 'Filter Particles...' button by size and average intensity; if you are done, press 'Select Markers'
- now markers can manually be postprocessed, i.e. markers can be added or removed, or their type can be changed
- at the end you can view marker statistics (button 'Results'), save the markers to a file (button 'Save Markers') or do some measurements (button 'Measurements...')
Functions and Options
Below we outline the functions of the various elements of the graphical user interface.
- Initialization:
- 'Initialize': initializes the plugin with the active image, if the image has more than one channel only the first channel is considered
- 'Keep original': if checked the source image remains open, otherwise it is closed
- Pre-segmentation:
- 'Detect': runs the particle detector
- 'Configure Operator...': allows to change the parameters of the particle detector
- 'Filter Particles...': allows to filter detection results
- 'Show contours': enables/disables display of the contours of detected particle regions
- 'Select markers': selects the final set of markers and terminates detection stage
Detected particles are labeled with marker type 1 and the counter of that type refers to their number.
- Manual post-processing:
- 'Add': add a new marker type at the end of the list, the type gets a random color
- 'Remove': delete the last type from the end of the list
- 'Delete': delete the last placed marker
- 'Reset': deletes all markers
- 'Show Markers': enables/disables display of markers
- 'Show Numbers': enables/disables display of marker numbers
- 'Show All': enables/disables display of both markers and numbers
- Results:
- 'Results': shows table with marker statistics
- 'Save Markers': saves the markers to an XML file
- 'Load Markers': loads markers from an XML file
- 'Export Image': save a copy of the image including all markers
For the pre-segmentation of spot-like structures a particle detector operator of MiToBo is used. The operator offers the following parameters:
- Parameters:
| Name | Description |
| Minimal Scale (JMin) | scale of smallest particles, integer value >= 1 |
| Maximal Scale (JMax) | scale of largest particles, integer value > JMin |
| Scale Interval Size | several adjacent scales are correlated, the interval size determines how many scales are considered |
| Correlation Threshold | threshold for detecting particles, the smaller the threshold is chosen the more particles will be detected |
| Minimum Region Size | minimal size of valid particles, smaller particles are discarded |
Sample data
Will be provided soon...