Applications/PaCeQuant: Difference between revisions
| Line 209: | Line 209: | ||
in the console. | in the console. | ||
The vignette contains comprehensive documentation of the various functions and | The vignette contains comprehensive documentation of the various functions and options of the package.<br> | ||
In addition, a sample workflow R script is provided which shows all commands to load your data, configure the output and do the analysis.<br> | In addition, a sample workflow R script is provided which shows all commands to load your data, configure the output and do the analysis.<br> | ||
You can directly copy the script into a file and load this file into the Rstudio editor to run it completely or line-wise. | You can directly copy the script into a file and load this file into the Rstudio editor to run it completely or line-wise. | ||
Revision as of 15:37, 21 December 2018
PaCeQuant Plugin

PaCeQuant is available since release version 1.8.6 of MiToBo.
The PaCeQuant plugin as part of MiToBo as well as the R script for the visualization of result data are published under the terms of GNU General Public License v3.0.


Related Publication
When using PaCeQuant, please cite us:
- B. Möller, Y. Poeschl, R. Plötner, and K. Bürstenbinder,
PaCeQuant: A Tool for High-Throughput Quantification of Pavement Cell Shape Characteristics.
Plant Physiology, 175(1), September 2017.
Name of Plugin/Operator
de.unihalle.informatik.MiToBo.apps.cellMorphology.PaCeQuant
(available since MiToBo version 1.8.6)
Main features
- extraction of 27 characteristic shape features to quantify pavement cell shape
- fully automatic segmentation of cell regions from input images
- optional import of external/manual cell segmentation data
- classification of lobes into type I (2-cell contact) and type II (3-cell contact)
- additional R scripts for feature visualization
Usage - Parameters
To run PaCeQuant perform the following steps:
- install MiToBo by following the instructions on the Installation page
- run MiToBo and start the operator runner by selecting the menu item PaCeQuant from Plugins -> MiToBo
This will bring up the operator window of PaCeQuant.
Phases and Operation modes
PaCeQuant supports three different options for running listed below:
- SEGMENTATION_AND_FEATURES:
expects images as input, segments the images and extracts features - SEGMENTATION_ONLY:
expects images as input, just segments the cell regions, no feature extraction - FEATURES_ONLY:
works on binary or label images or ImageJ regions, extracts features for the given regions
In addition PaCeQuant can be run in either of two modes:
- INTERACTIVE:
PaCeQuant processes the data provided directly within the graphical environment of ImageJ, i.e. reads regions from the ROI manager, and directly shows results - BATCH:
PaCeQuant processes all files (images or ROI files) in a given folder and writes results to disk
For batch mode the user can specify an input directory, in interactive mode PaCeQuant expects an input image or regions to be available in ImageJ.
Depending on the chosen phase and operation mode the graphical user interface is dynamically re-configured to show only the options relevant for the selected options.
Input Image/Directory
Depending on the chosen phases and operation modes here you need to specify either input data already loaded in ImageJ/Fiji or a directory where PaCeQuant can find the data to process. In detail, you have to provide the following information for the various configurations:
| Operation Mode | Phase(s) to Run | Input Data |
| INTERACTIVE | SEGMENTATION_ONLY | Gray-scale input image already opened in ImageJ/Fiji. |
| SEGMENTATION_AND_FEATURES | Gray-scale input image already opened in ImageJ/Fiji. | |
| FEATURES_ONLY | Binary or label image already opened in ImageJ/Fiji, or ImageJ ROI set from ROI manager. | |
| BATCH | SEGMENTATION_ONLY | Directory containing gray-scale input images. First-level sub-folders are also processed. |
| SEGMENTATION_AND_FEATURES | Directory containing gray-scale input images. First-level sub-folders are also processed. | |
| FEATURES_ONLY | Directory containing either binary or label images to process, a collection of ImageJ ROI files (with ending '.roi'), or an archive of multiple ImageJ ROIs (with ending '.zip'). |
Important notice:
In BATCH mode PaCeQuant tries to analyze all image files present in the given folder or any of its direct sub-folders. In particular, PaCeQuant will also analyze data from a potential result folder of a previous run, if it is present in the given folder. To avoid problems resulting from analyzing wrong data you should ensure that the provided directory only contains native input images or segmentation data and no other image data not suitable for processing with PaCeQuant.
Calibration
As PaCeQuant measures lengths and areas from the given data it is important that the tool is properly calibrated, i.e. the physical size of a pixel is known. PaCeQuant supports two calibration modes:
- AUTO:
here PaCeQuant seeks to extract calibration information from the given input data - MANUAL:
the user can enter calibration data, i.e. the physical size of a pixel and the units to use
Important notice:
Please note that in ImageJ ROI files no calibration data is stored. Thus, when using PaCeQuant to extract features from external segmentation data provided as ImageJ ROIs, you always need to manually provide calibration data. And this also holds when manually post-processing segmentation data extracted with PaCeQuant as all calibration data of the original images will get lost.
Configuration of Segmentation Phase
For detailed configuration of the algorithms applied during the segmentation phase the following parameters are available:
| Name | Description |
| Border Contrast | Allows to select whether the boundaries of the cells are darker or brighter than the background. |
| Heuristic for Gap Closing | During segmentation sometimes small gaps in the boundaries remain which can be closed by PaCeQuant applying one of the following heuristics:
|
| Unit for Size Thresholds | Unit in which the size thresholds for filtering valid regions (see next two parameters) are specified, i.e. either PIXELS or MICRONS. |
| Minimal Size of Cells | Segmented cells being too small can automatically be excluded by specifying a minimal size for valid cells. |
| Maximal Size of Cells | Segmented cells being too large can also automatically be excluded by specifying a maximal size for valid cells. |
Configuration of Feature Extraction Phase
| Name | Description |
| Feature Extraction | For changing various parameters used in feature extraction the MorphologyAnalyzer2D operator of MiToBo is used and can be configured here. Note that changing parameters might hamper comparison of PaCeQuant results among different work groups or laboratories. |
| Analyze lobe types? | Activates the optional classification of individual lobes into type I (2-cell contact) or type II (3-cell contact) lobes. |
Additional Configuration Parameters
| Name | Description |
| Draw region IDs to output image? | If the segmentation phase is run PaCeQuant outputs a label image showing segmented cell regions. If this option is activated region IDs are drawn to that image for easier interpretation. But note that this renders the image unsuitable for any further automatic analysis. |
| Verbose | If enabled additional log messages will be printed to console. |
| Show/save additional results? | If enabled an additional image stack with a collection of intermediate images will be generated. The images in this stack might help to get a deeper insight into PaCeQuant and might help to identify problems in case that the segmentation of input images fails. |
| Show/save feature stack? | If enabled a stack of images is generated where each image visualizes the values of a specific feature. For most images in this stack the feature values of individual cells are mapped to the intensity value of the corresponding cell (e.g., for features like area, solidity, width, length, branch count, etc.). |
Usage - Output data
If PaCeQuant is run in INTERACTIVE mode all results are directly shown in ImageJ/Fiji, while in BATCH mode all result data is saved to a folder named results in the input folder. Which kinds of output data are resulting from running PaCeQuant depends on the selected phases:
- list of result files from SEGMENTATION phase:
- *-allRois.zip: ImageJ ROI archive containing all ROIs extracted from a single image
- *_<number>.roi: ROI file containing only ROI with ID numberextracted from single image
- *-grayscale-result.tif: gray-scale label image of extracted cell regions
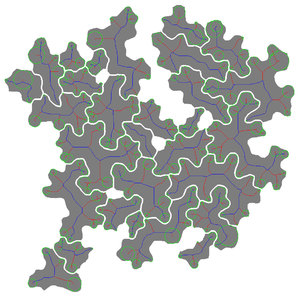
- *-color-result.tif: overlay of extracted cell regions (in pseudo-color) over input image
- *-intermediate-result-stack.tif: (optional) stack of intermediate result images
- list of result files from FEATURES phase:
- *-allRois-table.txt: extracted feature values for all cells in a given image
- *-allRois-grayscale-result.tif: gray-scale label image showing analyzed regions
- *-intermediate-result-stack.tif: (optional) stack of intermediate result images
- *-feature-stack.tif: (optional) stack of images visualizing feature values for cells
- *-allRois-lobe-types.tif: (if lobe type classification is enabled) image visualizing the type of each lobe of all cells analyzed
- *-allRois-cell-<number>-lobe-table.txt: (if lobe type classification is enabled) file containing features for lobes of cell with ID number
Note that the asterisk in the above file names represents a single image, i.e. the set of files exists for each image present in the given folder.
All files with ending *.ald are internal MiToBo log files in XML format where, e.g., parameter settings are stored and which usually can be ignored or just preserved for later reference.
Sample data
Here we provide some sample data with which you can test your local PaCeQuant installation:
To apply PaCeQuant to the test data simply unpack the data archive to a folder of your choice and set the input directory parameter of PaCeQuant to the sample data folder.
The second archive contains result data extracted with PaCeQuant for validating if your local PaCeQuant installation yields correct results.
PaCeQuantAna: R package for Visualization and Statistical Analysis of Cell Shape Featues
We provide an R package for visualizing and analyzing features extracted by PaCeQuant:
Installation
For installation in RStudio download the package and follow the instructions below:
- Install the following packages:
- caroline
- gplots
- dunn.test
- RColorBrewer
To this end start RStudio, select
from the menu bar.“Tools” → “Install Packages…”
Choose
and enter the names of the packages.“Install from: Repository (CRAN)”
Activate“Install dependencies”.
- The multtest package can be installed via the following Bioconductor installation script:
source ("http://bioconductor.org/biocLite.R")
biocLite("multtest")
- Install the PaCeQuantAna package selecting
“Tools” → “Install Packages…”
from the menu bar.
Choose
and select the tar.gz file of PaCeQuantAna via“Install from: Package Archive File (.tgz; .tar.gz)"
"Browse...".
Documentation
Find more details of how to use the package and details about the general workflow in the vignette of the package by either navigating to the vignette in the Rstudio package manager tab or by typing
vignette("PaCeQuantAna")
in the console.
The vignette contains comprehensive documentation of the various functions and options of the package.
In addition, a sample workflow R script is provided which shows all commands to load your data, configure the output and do the analysis.
You can directly copy the script into a file and load this file into the Rstudio editor to run it completely or line-wise.
You can test the package on these sample data: sample data for PaCeQuantAna
Important notes:
- By default the working directory (set with setpw(...)) where the result folder will be created, and the data directory (set with setDataDir(...)) must not be identical! If you require these folders to be identical for whatever reason you need to manually edit the file data_description.csv in the output folder and delete the row referring to the output folder.
Notes on manual or semi-automatic segmentation of cells
PaCeQuant supports extracting features from externally provided segmentation data.
This can be helpful in cases where the original input images are not suitable for automatic segmentation by PaCeQuant,
e.g., due to a general low image quality or image acquisition techniques for which PaCeQuant is not optimized.
The segmentation data can be provided in different formats:
- an 8-bit binary image where all cell regions are labeled white (intensity = 255)
- an 8-bit gray-scale image where each cell region is marked with an individual label (intensity larger than zero) and the background in black (intensity = 0)
- ImageJ ROI files
Note that when providing binary images neighboring regions must not touch each other. Optimally the margins between neighboring cell regions will have a width of at least 3 pixels.
ImageJ ROI files
For providing segmentation data in terms of ImageJ ROI files the easiest way to do this is to segment regions directly in ImageJ/Fiji.
Use the Polygon Section Tool for segmenting individual cells, add each of the polygons to the ROI manager, and finally save all ROIs to a file.
In case that you aim to improve an initial segmentation result of PaCeQuant, open the ROI files generated by PaCeQuant during the segmentation phase with ImageJ's ROI manager and edit the ROIs.
For editing polygon selections ImageJ offers a large variety of possibilities, e.g.:
- single points can be removed (Ctrl + click)
- new points can be added by splitting segments (Shift + click)
- the complete region can be moved
- the polygon can be interpolated, i.e. sub-sampled
More information on the various options can be found in the ImageJ documentation:
Probably of interest might also be the macro 'Modifying Polygon Selections'.